What's New
Monthly Newsletter
Don't miss the latest DSI news. Subscribe to our newsletter & read the latest volume.
Upcoming Seminar
Our seminar series is currently on a break. Contact DSI-Seminars [at] llnl.gov (DSI-Seminars[at]llnl[dot]gov) with any questions.
Data Scientist Spotlight

Vic Castillo
Computational Engineer
Computational engineer and data scientist Vic Castillo helps domestic manufacturers improve their processes—a challenge he finds rewarding. As a 30-year LLNL employee, Castillo has worn many hats, especially in support of Strategic Deterrence and Global Security programs. Castillo first came to the Lab as a graduate student through UC Davis’s Applied Science program (a.k.a. Teller Tech), having previously worked as a scientist at the Clorox Research Center. Castillo is now assisting the High-Performance Computing for Energy Innovation (HPC4EI) Program and has received 12 grants to collaborate with domestic manufacturing companies. He says that analyzing real-world data can be laborious, but HPC simulation and machine learning tools have vast potential to improve manufacturing processes and reduce carbon emissions. “My work feels like a small consulting business with many clients,” he states. “There are many different types of problems, and the Lab has broad capabilities to meet these challenges.” Castillo frequently speaks to manufacturing and university audiences to publicize the Lab’s capabilities and publishes journal articles with his team. He has mentored 25 students and postdocs during his career and helped to create an LLNL Teacher Research Academy for local STEM teachers.
Recent Research
ML Tool Fills in the Blanks for Satellite Light Curves
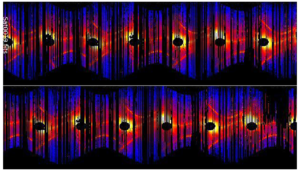
When viewed from Earth, objects in space are seen at a specific brightness, called apparent magnitude. Over time, ground-based telescopes can track a specific object’s change in brightness. This time-dependent magnitude variation is known as an object’s light curve, and can allow astronomers to infer the object’s size, shape, material, location, and more. Monitoring the light curve of satellites or debris orbiting the earth can help identify changes or anomalies in these bodies. However, light curves are missing a lot of data points. The weather, the season, dust accumulation, time of day, eclipses—these all affect not only the quality of the data, but whether it can be taken at all. Livermore researchers have developed an ML process for light curve modeling and prediction. Called MuyGPs, the process drastically reduces the size of a conventional Gaussian process problem by limiting the correlation of predictions to their nearest neighboring data points, reducing a large linear algebra problem to many smaller, parallelizable problems. This type of ML enables training on more sensitive parameters, optimizing the efficient prediction of the missing data. Read more about MuyGPs.