New technological advances and the cheapening of data acquisition have vastly expanded what is possible in bioinformatics. Things like predicting protein folding and interactions, which I previously believed impossible, are not anymore. My experience at LLNL has changed what I think is possible.
—Jonathan Anzules, DSSI class of 2022
As a DSSI intern, you’ll work on real projects with real data that represent the breadth and depth of data science research at LLNL. Our students tackle Challenge Problems that leverage large and varied datasets used in or generated from actual LLNL projects such as building networks from interaction data, large-scale data mining for predictive medicine, drug discovery using HPC simulations, video data summarization and classification, energy efficiency analysis using HPC, classification and forward modeling of hyper-spectral data, and much more. This curriculum helps students build technical experience and teamwork.
Our interns are also paired with mentors—experts across many data science fields at the Lab—whose projects align with students’ skills and interests. Check out recent Challenge Problems and mentors below.
2023 Challenge Problem
Mentors: Mikel Landajuela and team
A highly useful and commonly used diagnostic tool, the electrocardiogram (ECG) provides a noninvasive, cost-effective diagnosis of heart conditions. But the standard 12-lead ECG is inadequate for mapping the heart’s electrical activity in sufficient detail for many clinical applications.
The students' tasks ranged from a simple classification problem—using machine learning to distinguish a healthy heart from an abnormal heart and diagnosing the condition—to the most complicated task of reconstructing a full heart activation map from 12-lead ECG data taken from 75 areas of the heart. The models can be used for heartbeat simulations and more advanced diagnostics of heart conditions.
Challenge: Explore a data-driven approach to reconstructing electro-anatomical maps of the heart at clinically relevant resolutions, combining input from the standard 12-lead ECG with advanced machine learning techniques.
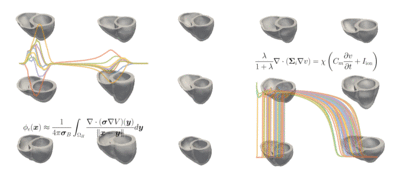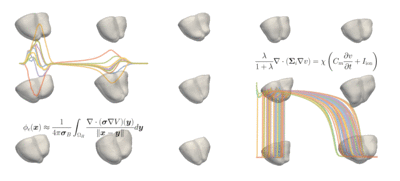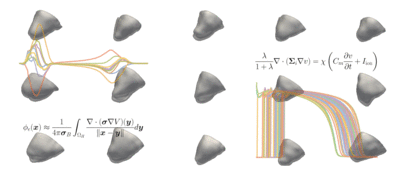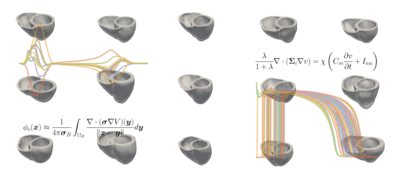
Read more about Machine Learning for Cardiac Electrocardiography and download the code on GitHub.
2022 Challenge Problem
Mentors: Hyojin Kim, Garrett Stevenson, and team
Since the beginning of the COVID-19 pandemic, the world has seen more than 600 million cases of the virus and more than 6 million deaths. LLNL is actively engaged in developing medical countermeasures for this and other emerging pathogens. Traditional drug discovery involves many time-consuming and expensive experimental steps, and unpredictable variants can complicate the process. Machine learning and other data science techniques can drastically accelerate drug discovery, which is especially important in a global pandemic.
For instance, LLNL researchers have been computationally screening hundreds of millions of small-molecule inhibitors to identify a subset that can be developed into antiviral drugs for COVID-19. When potential candidates are identified, we conduct additional machine learning screens for improving safety and pharmacokinetic profiles of the potential drugs.
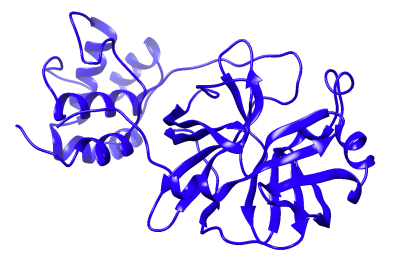
Challenge: Work with datasets of virtual molecule screening results, chemical and protein structures, and designed synthetic antibodies to identify drug compounds that can be used to create medicines that prevent and treat COVID-19 infections. This means developing machine learning approaches that find small-molecule viral inhibitors with the potential to bind to the main protease receptors of SARS-CoV-2 (the virus that causes COVID-19).
Tasks to predict binding affinity:
- Use molecular descriptors to predict MM/GBSA (molecular mechanics/generalized born surface area) values that correlate with experimental binding affinity
- Train a 3D convolutional neural network (CNN) to predict binding affinity between protein receptors and ligands
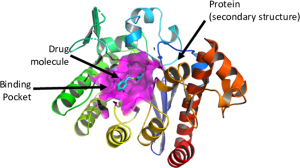
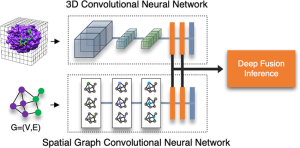
COVID-19 case data from the World Health Organization. Images from Jones D., Kim H., Zhang X., et al. (2021). “Improved Protein−Ligand Binding Affinity Prediction with Structure-Based Deep Fusion Inference.” Journal of Chemical Information and Modeling.
2021 Challenge Problems
Mentors: Ryan Dana, Kerianne Pruett, and team
Understanding the nature of dark matter and dark energy relies on accurately mapping the universe around us, making distinguishing stars from galaxies a crucial task for astronomers. If correctly identified, star and galaxy images can help researchers unravel the mysteries of the universe. With ever-increasing large-scale photometric surveys that collect data on upwards of billions of stars and galaxies, automated classification solutions have a significant time-saving advantage over manual classification methods.
Galactic objects emit waves on the electromagnetic spectrum. Stars and galaxies emit in multiple wavelength ranges. The Hyper Suprime-Cam (HSC) telescope in Hawaii uses photometric bands that allow in light from certain ranges. Its camera measures the amount of light that falls on each pixel during exposure.
Challenge: Use data science techniques to identify stars and galaxies in images from a ground-based telescope. Each image in a publicly available dataset of 34,000 images from the HSC consists of a 26x26 pixel array.
Tasks:
- Preprocess and normalize the data to increase accuracy and decrease runtime
- Build an image classifier to classify stars and galaxies
- Train the classifier on labeled images
- Report relevant statistics in accuracy
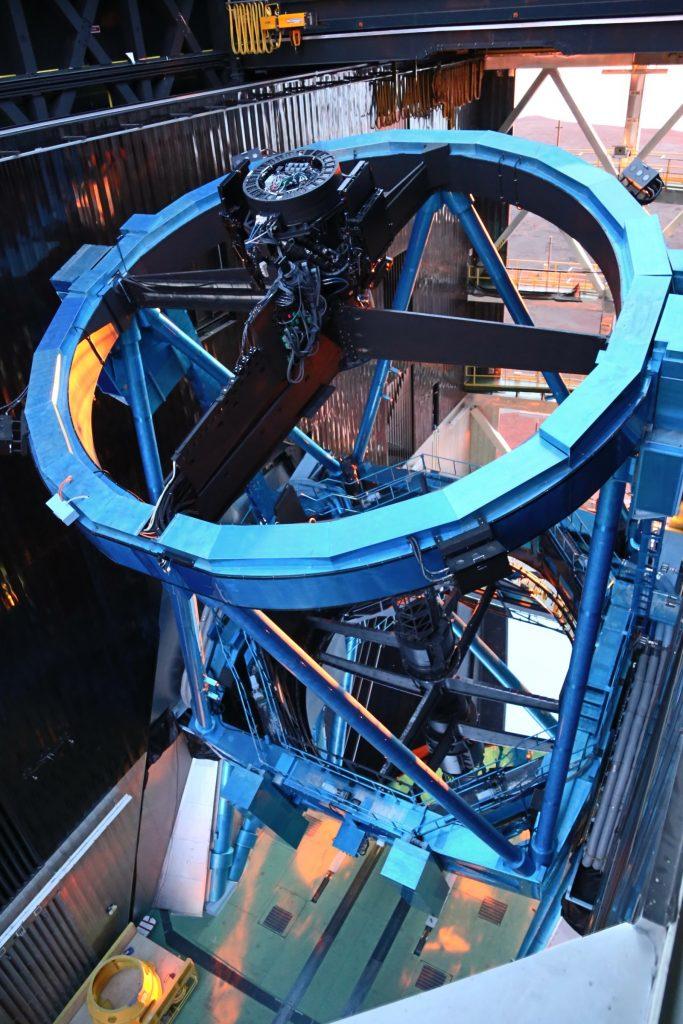

Part of the HSC telescope assembly. (Photo from https://hsc.mtk.nao.ac.jp/ssp/gallery/. Click to enlarge.)
Mentors: Ryan Dana, Kerianne Pruett, and team
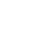
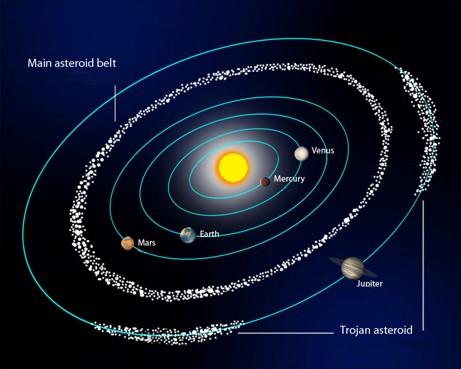
Smart planetary defense strategies require space situational awareness. Of the millions of orbiting asteroids and comets in the Solar System, several thousand are potentially hazardous to Earth. Larger objects are easier to detect but could inflict more damage. Finding these objects within telescope images is difficult because they reflect sunlight instead of emitting their own light—but brightness alone does not correspond to size.
The convolutional neural network (CNN) is a class of deep learning neural networks. CNNs represent a major breakthrough in image recognition, are most commonly used to analyze visual imagery, and are frequently working behind the scenes in image classification. Gaussian processes and Bayesian classifiers are other methods that may be used in this application.
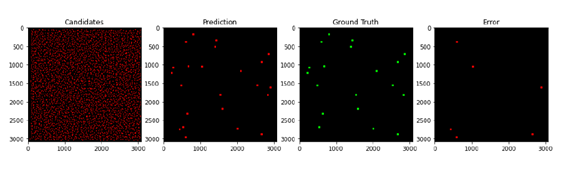
Challenge: Use deep learning techniques to identify asteroids. This smaller dataset of difference 1,000 images from California’s Zwicky Transient Facility (ZTF) have been injected with 20 asteroids each, for a sample size of 20,000 asteroids.
Tasks:
- Build an image classifier to detect asteroids
- Apply detection algorithms to the dataset to determine asteroid orbits and other characteristics
- Report relevant statistics in accuracy
Mentor Spotlight

Hyojin Kim
Hyojin Kim is a data scientist and machine learning researcher at LLNL’s Center for Applied Scientific Computing. His research interests in machine learning and computer vision are recently related to applications for computed tomography, AI-driven drug discovery, scalable and distributed deep learning, and multimodal image analysis. He also has hands-on experience applying GPU computing to challenging problems in these areas. Balancing research and development, as well as learning domain knowledge, are crucial because, Kim says, “I often see data scientists trying to apply a new technique to a particular domain application where it may not be suitable.” This summer, Kim mentored students from two University of California campuses in DSI’s Data Science Challenge to accelerate drug discovery for COVID-19. During the intensive two-week program, he states, “Many of the students I met were enthusiastic, and some of them came up with brilliant ideas that I never thought about before. Students majoring in fields other than computer science are quite knowledgeable in data science, and I actually feel the growing popularity of data science in recent years.” Kim joined LLNL in 2013 after earning his Ph.D. in Computer Science from UC Davis in 2012.
Ready for the next step?
Visit our applications page to read FAQ about the program, apply to the job posting, or contact us for more information. Join the LLNL Scholars Google Group to chat with mentors and other students.