- Home
- Research Areas
- Research Archive: Molecular Markers


"We look for key patterns such as molecular signatures that correlate with certain outcomes to build a model of the drugs’ effectiveness at countering tumor growth.”
– Jonathan Allen
The amount of experimental molecular measurements, such as genomes and gene expression, is growing at a rapid rate. Access to this data presents new opportunities to learn more of the molecular drivers of disease and to develop improved options for treatment. LLNL bioinformatics scientist Jonathan Allen leads a team exploring the use of both data-driven predictive models from experimental data as well as more efficient genome search and retrieval algorithms to design a suite of tools for pathogen detection and countermeasure design. Examples of questions the team is pursuing include finding unique genetic markers indicative of antibiotic resistance as well as modeling and predicting response to drug treatments for cancer. High-performance computing is a valuable tool in these efforts, pushing the limits on the size of data sets used to identify important functional molecular features.
Team Acknowledgments
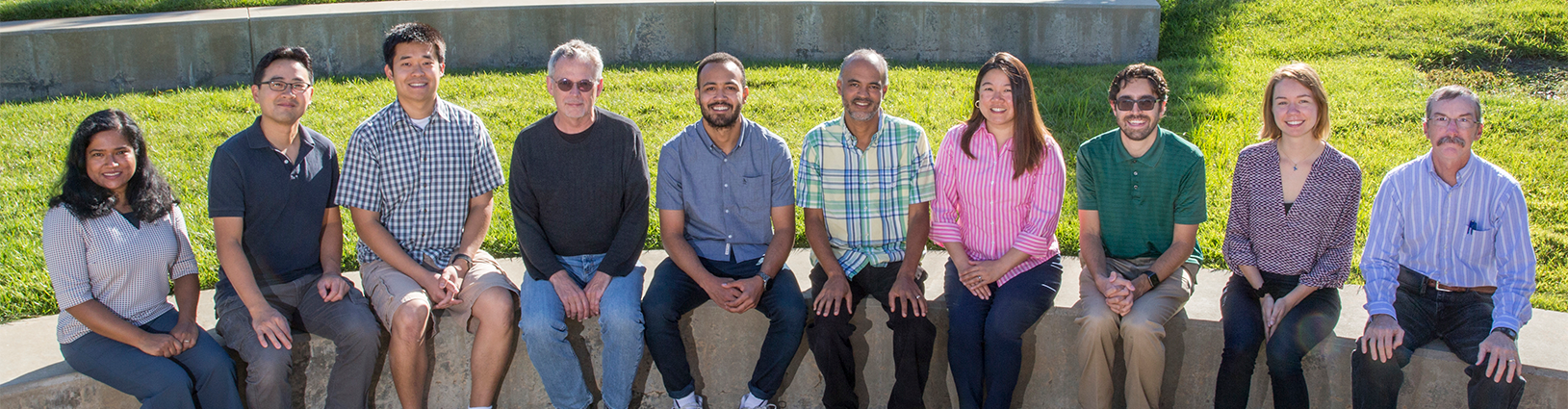
Pictured (left to right): Nisha Mulakken, Hyojin Kim, Stewart He, Kevin McLoughlin, William Jones, Jonathan Allen, Marisa Torres, Ryan Forsyth, Masha Aseeva, and Mark Wagner.
Not pictured: Aram Avila-Herrera, Aiden Epstein, Ya Ju Fan, Dayanara Lebron Aldea, Amanda Minnich, and Adam Zemla.