Feb. 24, 2022

Register for Virtual WiDS Livermore on March 7
The annual Women in Data Science (WiDS) conference returns on Monday, March 7, which is International Women’s Day. LLNL will again host a regional event in conjunction with the worldwide conference. The all-day WiDS Livermore event will be entirely virtual and free. Everyone is welcome to attend. Registration is open until February 27.
Sponsored by the DSI and LLNL’s Office of Strategic Diversity and Inclusion Programs, WiDS Livermore will include a livestream of the Stanford conference and networking opportunities. Returning this year is the popular “speed mentoring” session, where mentees rotate through breakout rooms to speak with mentors. Attendees can sign up to be mentors or mentees during registration. To mark LLNL’s fifth year of WiDS participation, the Livermore event will feature a fireside chat between Kim Budil, the Lab’s first woman director, and Jessie Gaylord, leader of Computing’s Global Security Computing Applications Division. Afternoon workshops will give participants a chance to learn the basics of data science or apply their skills. Gale Lucas from the University of Southern California’s Institute for Creative Technologies will lead a Beginner workshop track, while LLNL’s Gemma Anderson will explore machine learning for climate predictions in the Applied workshop track. The Applied track will also include a presentation by LLNL data scientists Olivia Miano and Juanita Ordoñez.
Winter Hackathon Meets WiDS Datathon
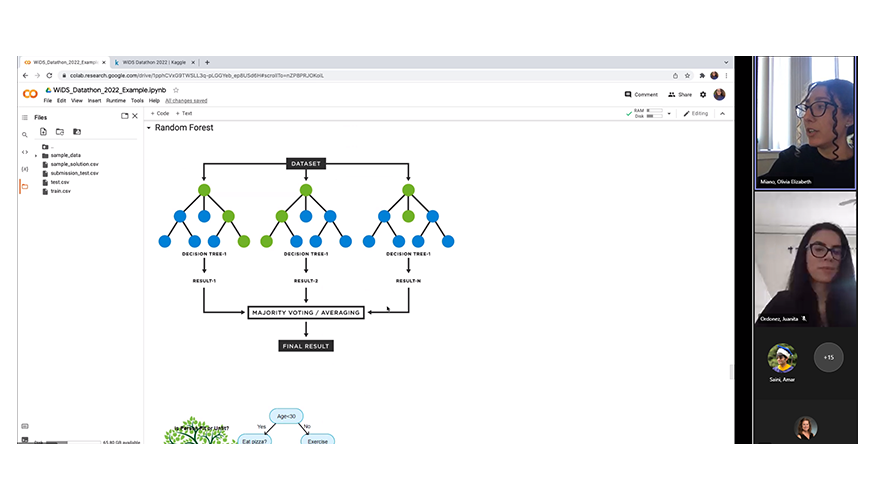
Sponsored by the DSI, LLNL’s winter hackathon took place on February 16–17. Hackathons are 24-hour events that encourage collaborative programming and creative problem solving. In addition to traditional hacking, the hackathon included a special datathon competition in conjunction with the upcoming WiDS Livermore event (see previous story). The datathon challenged participants to create a predictive model using a climate dataset curated by the global WiDS organization, and teams had the option of submitting their results to the global competition.
LLNL data scientists Olivia Miano and Juanita Ordoñez provided participants with a Jupyter Notebooks tutorial on how to analyze the datathon dataset with Pandas. They demonstrated applying various machine learning models to the data, including linear regression, root mean squared error, random forest (shown at left), and gradient boosting regression.
Continuing the datathon theme, LLNL researchers Gemma Anderson and Aaron Donahue presented an overview of global climate models, explaining how the Lab uses deep learning to extract complex patterns and connections from climate data.
Hackathon and datathon participants presented their projects on the second day. LLNL event organizers were Cindy Gonzales, Amar Saini, Mary Silva, and Jennifer Bellig.

Improving Deep Learning for Materials Optimization Analysis
Although many deep learning (DL) model explanation methods exist for image-based input, they are tailored for natural images and do not translate well to scientific image prediction tasks. What matters in scientific images is usually not a few distinctive objects but the distribution of objects and the interaction between different objects.
An LLNL research team has published an approach in ACS Omega leveraging a generative adversarial network (GAN)—a DL method that generates high-quality images—that modifies images based on human interpretable domain attributes for model explanation. This GAN model successfully injects domain attributes in a post hoc manner into the prediction pipeline with only a few data labels, allowing researchers to explain DL decision making in the language that a domain scientist can understand. For example: How does a change in the crystal size (or porosity, etc.) impact the peak stress prediction? How should material attributes be altered to obtain a material with higher peak stress? The research team includes Shusen Liu, Bhavya Kailkhura, Jize Zhang, Anna Hiszpanski, Emily Robertson, Donald Loveland, Xiaoting Zhong, and Yong Han.
(Image at left: Illustration of material attributes–guided scanning electron microscope image generation. The left column shows the original image. The middle and right columns show the GAN-generated images of hypothetical lots that increase or decrease the corresponding material attributes, respectively. The colored boxes highlight the corresponding regions where changes reflect the alterations in the attribution.)
Dynamic Model of COVID-19 Disease Progression
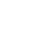
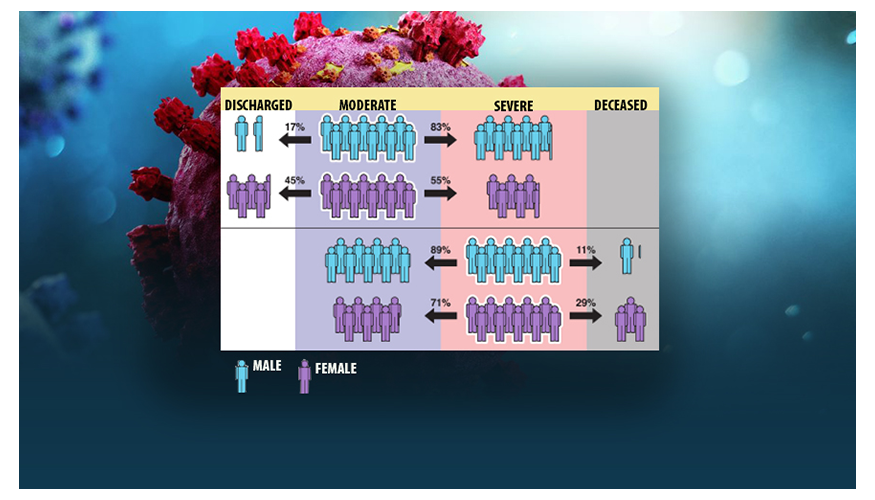
Being male is a known risk factor for adverse outcomes in hospitalized COVID-19 patients. However, new analysis reveals that when modeling the entire disease trajectory, the degree to which being male is a risk factor depends on the underlying disease severity of the patient. An LLNL team has developed a comprehensive dynamic model of COVID-19 disease progression in hospitalized patients, finding that risk factors for complications from the disease are dependent on the patient’s disease state.
Using a machine learning algorithm on a dataset of electronic health records from more than 1,300 hospitalized COVID-19 patients with ProMedica, the team classified patients into “moderate” or “severe” states and tracked disease trajectory as patients moved through different risk states during hospitalization. Accounting for disease severity—in contrast to previous scientific literature examining only static risk factors—the method allowed the team to identify, as the disease progressed, when certain variables such as age and race, and comorbidities including diabetes and hypertension, led to more severe outcomes.
The results are published in the Journal of the American Medical Informatics Association. Co-authors are Braden Soper, Jose Cadena, Sam Nguyen, Kwan Ho Ryan Chan, and Priyadip Ray with colleagues from ProMedica Health System and the University of Toledo College of Medicine and Life Sciences.

Virtual Seminar Explores Data-Driven Mechanistic Models
For the DSI’s January 20 virtual seminar, Dr. Babak Shahbaba of UC Irvine presented “Data-Driven Mechanistic Models – Design Inference.” Using a Bayesian framework, he focused on the application of mechanistic models for investigating dynamic biological systems. Shahbaba also proposed a class of scalable Bayesian inference methods that utilize deep learning algorithms for fast approximation of the likelihood function and its gradient.
Dr. Shahbaba’s research focuses on Bayesian methods and their applications in data-intensive biomedical problems. His experience spans a broad spectrum of areas including statistical methodologies (Bayesian nonparametrics and hierarchical Bayesian models), computational techniques (efficient sampling algorithms) and a wide range of applied and collaborative projects (statistical methods in neuroscience, genomics, and health sciences). A recording of his talk will be posted to the seminar series’ YouTube playlist

Meet an LLNL Data Scientist
Ryan Dana is a data scientist working in the Global Security Computing Applications Division and the astrophysical analytics group. He was a student intern with LLNL’s Data Science Summer Institute (DSSI) in 2019 and graduated that year with a B.A. in Physics, Astrophysics, and Data Science from UC Berkeley. “The DSSI perfectly connected my coursework in physics, astronomy, statistics, and computer science to solving problems using real-world astronomical data,” he says. Dana joined the Lab as a full-time employee in early 2020 and mentored students in the 2021 Data Science Challenge program—a position he returns to in 2022. His research interests include using machine learning techniques to approach astrophysical questions.

UC Merced Data Science Challenge Coming Soon
This month UC Merced students began applying for the next Data Science Challenge. Now in its fourth year, the program is a close collaboration with the university, giving undergraduate and grad students the opportunity to grow their skills while exploring a real-world scientific problem. This year, the students will work with a database of virtual molecule screening results, chemical and protein structures, and designed synthetic antibodies to identify drug compounds that can be used to create medicines that prevent and treat COVID-19 infections. Led by LLNL’s Brian Gallagher and UC Merced mathematics professor Suzanne Sindi, the program’s mentors include LLNL researchers Hyojin Kim, Ryan Dana, and Cindy Gonzales. Students have until March 13 to apply, and the Challenge will run from May 23 to June 6.